YMLA Tutorial
YMLA focuses on performing comparative enrichment analysis for multiple gene lists in yeast.
1. Input page
3. Heatmaps, and Network Views
5. Detail page
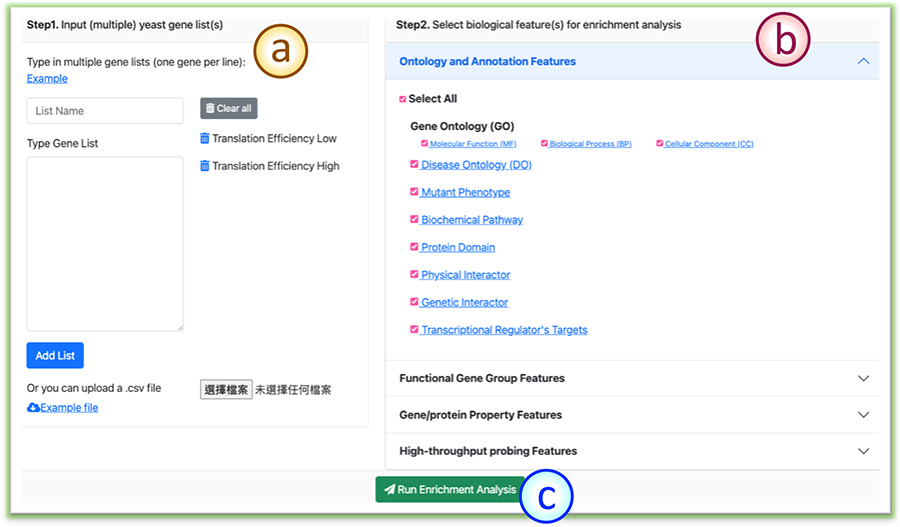
(a) Users can provide the gene lists of interest either via the text area or a .csv file.
(b) Users also need to choose the features to be included in the analysis.
(c) Click the ”Run Enrichment Analysis” button to begin the evaluation.
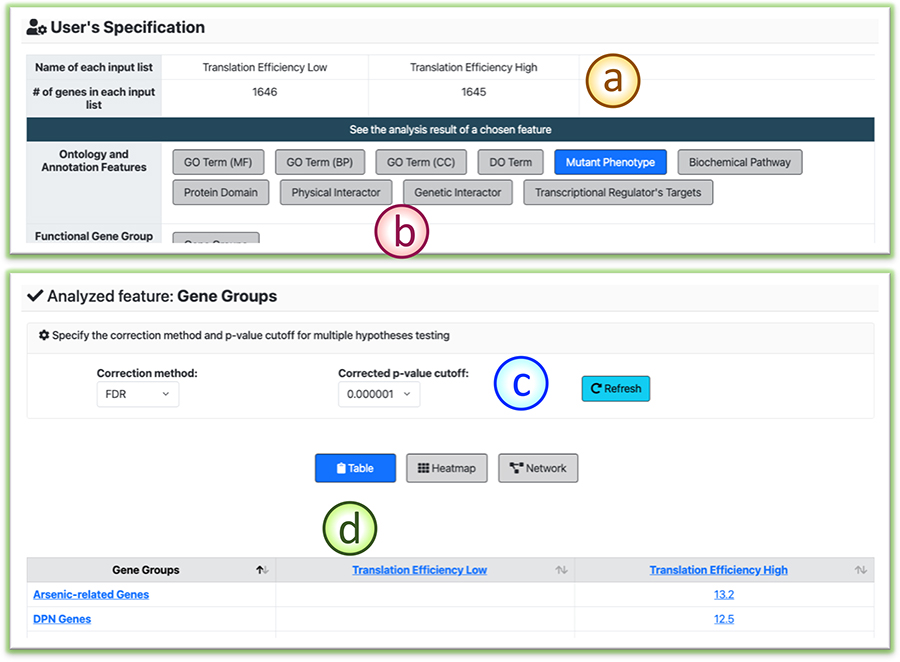
(a) The gene item summary for the input multiple gene lists.
(b) Users can select the feature of interest to investigate the enrichment analysis results.
(c) Users can reset the multiple hypotheses correction methods and the significance thresholds.
(d) The tabular view of the enrichment analysis results. Users can click the hyperlinks to navigate the detailed results.
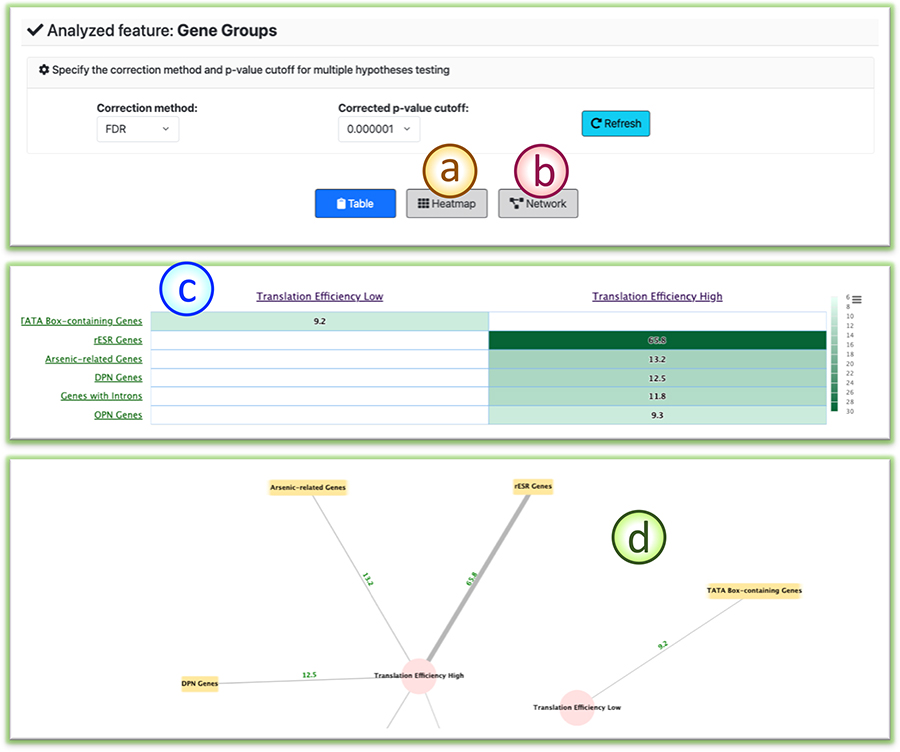
(a)/(b) Users can click the tab to switch to the heatmap/network view of the analysis results.
(c) The heat map view of the analysis results.
(d) The network view of the enrichment analysis results.
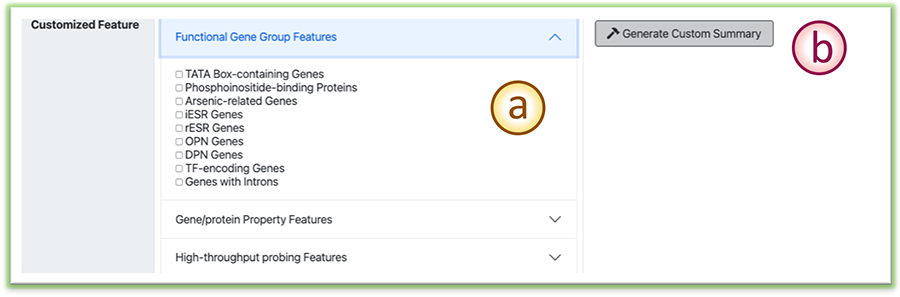
(a) Users can pick the features related to some mechanisms hypotheses to form a new enrichment analysis.
(b) Click the ”Generate Custom summary” button to re-perform the enrichment analysis using the customized feature set.
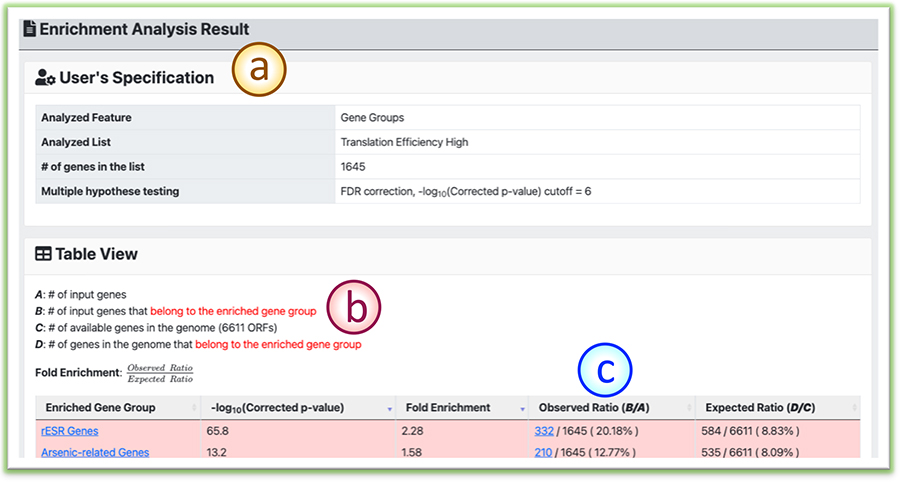
When the hyperlinks in the analysis result tables are clicked, the detailed page that supports the analysis will be shown.
(a) The item summary reminder of this detail page for users.
(b) Indication of how the enrichment scores are calculated in YMLA.
(c) The detailed enrichment results that support the presented enrichment analysis.